Solving the Poisson equation#
Solving 1D Poisson with periodic boundary conditions in Struphy#
This tutorial shows how to solve a manufactured 1D Poisson problem with the Poisson model.
We use a periodic domain \(x \in [0, L)\) and choose an analytic solution
so the source for the stabilized Poisson problem
is
Then we run a Struphy Simulation, compare numerical and exact solutions, and compute the max-norm error.
[1]:
import numpy as np
import matplotlib.pyplot as plt
from struphy import Simulation, DerhamOptions, domains, grids, perturbations
from struphy.initial.base import GenericPerturbation
from struphy.models import Poisson
Poisson.pde()
PDEs solved by model:
Find φ ∈ H¹ such that
where n₀, ρ(t) : Ω → ℝ are real-valued functions, ρ(t) is
parametrized by time t, and D₀ : Ω → ℝ3 × 3 is a positive matrix.
Boundary terms from integration by parts are assumed to vanish.
[2]:
# Manufactured periodic test case in 1D (aligned with verification test pattern)
Lx = 2.0 * np.pi
mode = 2
k = mode * 2.0 * np.pi / Lx
# Tiny stabilization makes the periodic problem uniquely solvable
stab_eps = 1e-8
# Build the source through the model variable as in test_verif_Poisson.py
model = Poisson()
model.propagators.poisson.options = model.propagators.poisson.Options(
rho=model.em_fields.source,
stab_eps=stab_eps,
)
source_amp = k**2 + stab_eps
model.em_fields.source.add_perturbation(
perturbations.ModesCos(ls=(mode,), amps=(source_amp,))
)
phi_exact = lambda e1, e2, e3: np.cos(k * e1)
Poisson.Options controls the elliptic solve. In this example we only set:
rho: the manufactured source term.stab_eps: a tiny stabilization to remove the constant null-space in the periodic case.
All other option fields are left at their defaults (solver choice, preconditioner, tolerances, etc.).
[3]:
# 1D periodic setup: periodic bcs are the default (None in each direction)
domain = domains.Cuboid(l1=0.0, r1=Lx)
grid = grids.TensorProductGrid(num_elements=(64, 1, 1))
derham_opts = DerhamOptions()
sim = Simulation(
model=model,
domain=domain,
grid=grid,
derham_opts=derham_opts,
verbose=False,
)
# For a stationary Poisson solve, one step is enough
sim.run(one_time_step=True, verbose=False)
sim.pproc(verbose=False)
sim.load_plotting_data(verbose=False)
Starting simulation run for model Poisson ...
Struphy run finished.
100%|██████████| 2/2 [00:00<00:00, 1220.34it/s]
100%|██████████| 2/2 [00:00<00:00, 67.17it/s]
100%|██████████| 2/2 [00:00<00:00, 1691.93it/s]
[4]:
# Extract 1D line data and compare to analytic solution
x = sim.grids_phy[0][:, 0, 0]
t_last = max(sim.spline_values.em_fields.phi_log.data.keys())
phi_num = sim.spline_values.em_fields.phi_log.data[t_last][0][:, 0, 0]
phi_ref = phi_exact(x, 0.0, 0.0)
err = phi_num - phi_ref
err_max = np.max(np.abs(err))
plt.figure(figsize=(9, 4))
plt.subplot(1, 2, 1)
plt.plot(x, phi_ref, "k--", label="analytic")
plt.plot(x, phi_num, "o", ms=3, label="Struphy")
plt.xlabel("x")
plt.ylabel(r"$\phi(x)$")
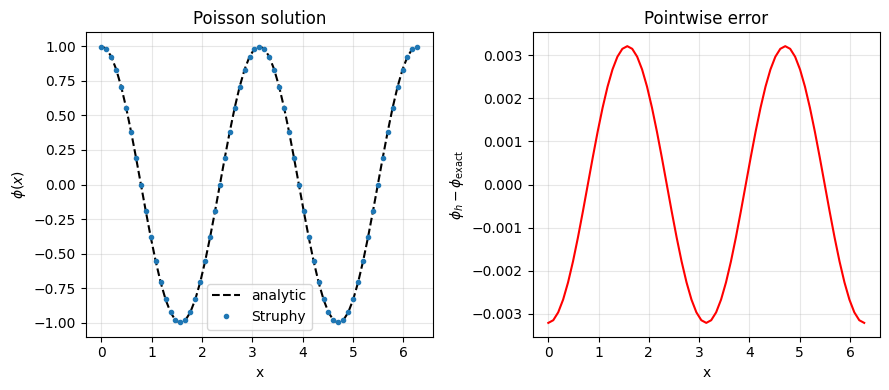
plt.title("Poisson solution")
plt.legend()
plt.grid(alpha=0.3)
plt.subplot(1, 2, 2)
plt.plot(x, err, "r", lw=1.5)
plt.xlabel("x")
plt.ylabel(r"$\phi_h - \phi_\mathrm{exact}$")
plt.title("Pointwise error")
plt.grid(alpha=0.3)
plt.tight_layout()
print(f"max-norm error ||phi_h - phi_exact||_inf = {err_max:.3e}")
# Electric field comparison: E = -grad(phi)
E_num = -np.gradient(phi_num, x, edge_order=2)
E_ref = k * np.sin(k * x)
E_err = E_num - E_ref
E_err_max = np.max(np.abs(E_err))
plt.figure(figsize=(9, 4))
plt.subplot(1, 2, 1)
plt.plot(x, E_ref, "k--", label=r"analytic $-\nabla\phi$")
plt.plot(x, E_num, "o", ms=3, label=r"Struphy $-\nabla\phi_h$")
plt.xlabel("x")
plt.ylabel(r"$E_x(x)$")
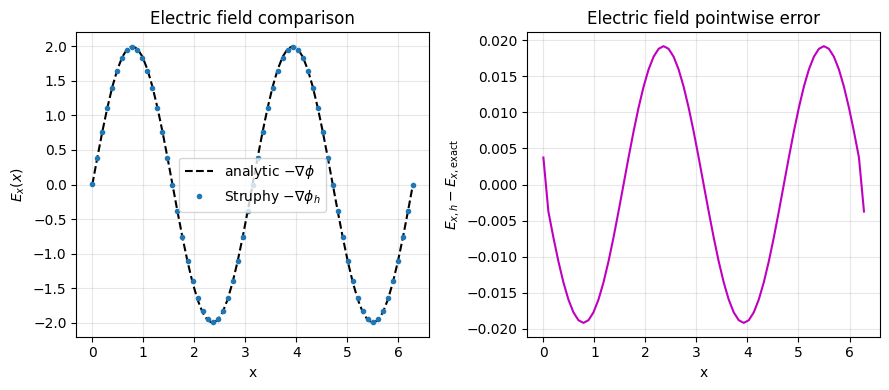
plt.title("Electric field comparison")
plt.legend()
plt.grid(alpha=0.3)
plt.subplot(1, 2, 2)
plt.plot(x, E_err, "m", lw=1.5)
plt.xlabel("x")
plt.ylabel(r"$E_{x,h} - E_{x,\mathrm{exact}}$")
plt.title("Electric field pointwise error")
plt.grid(alpha=0.3)
plt.tight_layout()
print(f"max-norm error ||E_h - E_exact||_inf = {E_err_max:.3e}")
max-norm error ||phi_h - phi_exact||_inf = 3.207e-03
max-norm error ||E_h - E_exact||_inf = 1.920e-02
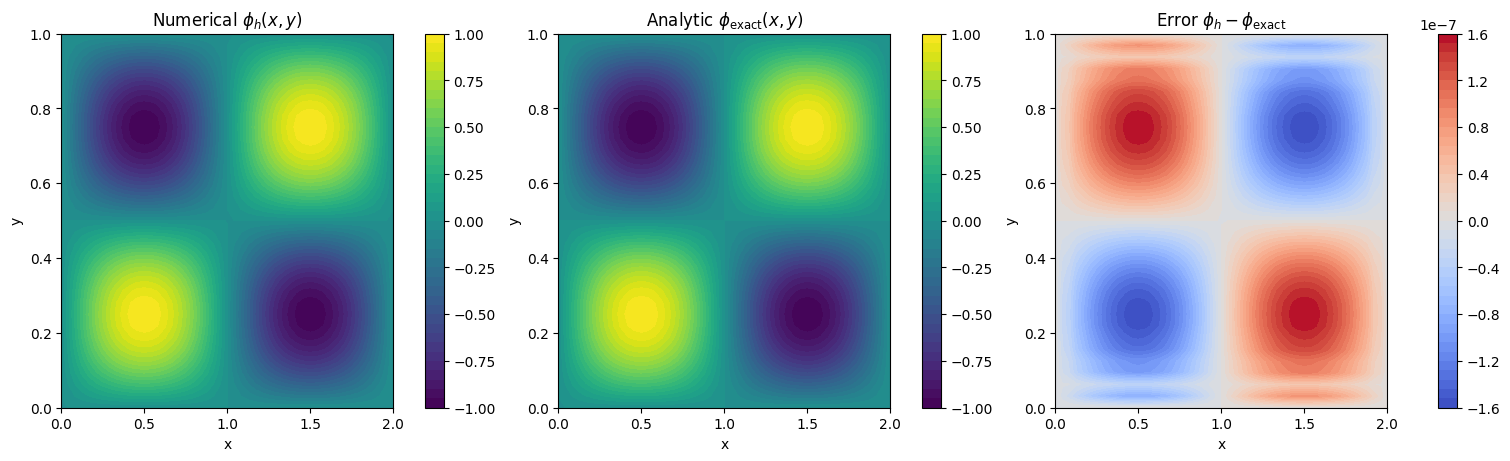
2D Manufactured Test on a Rectangle#
We now solve a 2D manufactured Poisson problem on a rectangle with different side lengths:
Choose
which satisfies homogeneous Dirichlet conditions on all rectangle boundaries. For
the manufactured source is
To stay consistent with Struphy’s source handling, we build this source through a model perturbation.
[5]:
# 2D manufactured Poisson setup and solve
Lx2 = 2.0
Ly2 = 1.0
eps2 = 1e-8
kx2 = 2.0 * np.pi / Lx2
ky2 = 2.0 * np.pi / Ly2
phi2_exact = lambda e1, e2, e3: np.sin(kx2 * e1) * np.sin(ky2 * e2)
model2 = Poisson()
model2.propagators.poisson.options = model2.propagators.poisson.Options(
rho=model2.em_fields.source,
stab_eps=eps2,
)
source_amp2 = kx2**2 + ky2**2 + eps2
model2.em_fields.source.add_perturbation(
perturbations.ModesSinSin(ls=(1,), ms=(1,), amps=(source_amp2,))
)
domain2 = domains.Cuboid(l1=0.0, r1=Lx2, l2=0.0, r2=Ly2)
grid2 = grids.TensorProductGrid(num_elements=(48, 32, 1))
derham_opts2 = DerhamOptions(
degree=(3, 3, 1),
bcs=(("dirichlet", "dirichlet"), ("dirichlet", "dirichlet"), None),
)
sim2 = Simulation(
model=model2,
domain=domain2,
grid=grid2,
derham_opts=derham_opts2,
verbose=False,
)
sim2.run(one_time_step=True, verbose=False)
sim2.pproc(verbose=False)
sim2.load_plotting_data(verbose=False)
Starting simulation run for model Poisson ...
Struphy run finished.
100%|██████████| 2/2 [00:00<00:00, 1321.25it/s]
100%|██████████| 2/2 [00:00<00:00, 2.76it/s]
100%|██████████| 2/2 [00:00<00:00, 264.65it/s]
[6]:
# 2D diagnostics and plots
t2_last = max(sim2.spline_values.em_fields.phi_log.data.keys())
X = sim2.grids_phy[0][:, :, 0]
Y = sim2.grids_phy[1][:, :, 0]
phi2_num = sim2.spline_values.em_fields.phi_log.data[t2_last][0][:, :, 0]
phi2_ref = phi2_exact(X, Y, 0.0)
err2 = phi2_num - phi2_ref
err2_max = np.max(np.abs(err2))
fig, axs = plt.subplots(1, 3, figsize=(15, 4.5), constrained_layout=True)
im0 = axs[0].contourf(X, Y, phi2_num, levels=40, cmap="viridis")
axs[0].set_title("Numerical $\\phi_h(x,y)$")
axs[0].set_xlabel("x")
axs[0].set_ylabel("y")
plt.colorbar(im0, ax=axs[0])
im1 = axs[1].contourf(X, Y, phi2_ref, levels=40, cmap="viridis")
axs[1].set_title("Analytic $\\phi_{\\mathrm{exact}}(x,y)$")
axs[1].set_xlabel("x")
axs[1].set_ylabel("y")
plt.colorbar(im1, ax=axs[1])
im2 = axs[2].contourf(X, Y, err2, levels=40, cmap="coolwarm")
axs[2].set_title("Error $\\phi_h-\\phi_{\\mathrm{exact}}$")
axs[2].set_xlabel("x")
axs[2].set_ylabel("y")
plt.colorbar(im2, ax=axs[2])
print(f"2D max-norm error ||phi_h - phi_exact||_inf = {err2_max:.3e}")
2D max-norm error ||phi_h - phi_exact||_inf = 1.589e-07
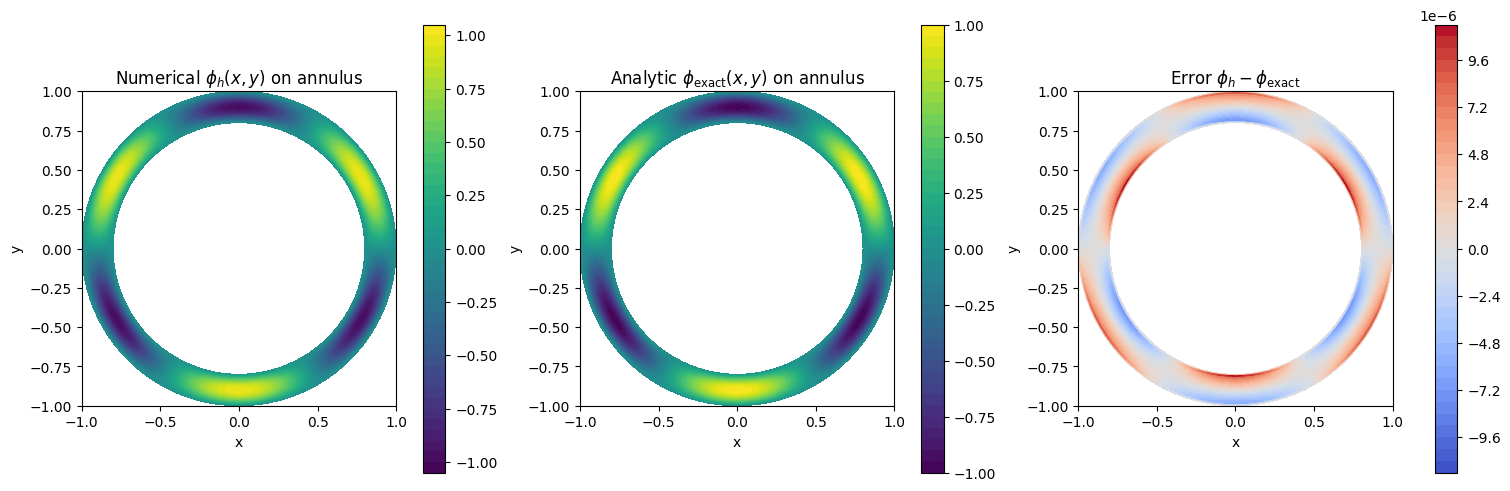
2D Manufactured Test on a Thin Annulus (HollowCylinder)#
As a second 2D geometry test, we solve Poisson on a disc-shaped domain with a small hole, i.e. a thin annulus.
We use the physical manufactured solution
with \(r=\sqrt{x^2+y^2}\) and \(\theta=\mathrm{atan2}(y,x)\). This vanishes at \(r=a_1\) and \(r=a_2\), so we impose homogeneous Dirichlet conditions in the radial direction.
For
the source \(\rho\) is computed analytically in polar coordinates and injected as a physical-space perturbation.
Below, all plots are shown in physical coordinates \((x,y)\) only.
[7]:
# HollowCylinder manufactured annulus test (physical-space definition)
a1 = 0.8
a2 = 1.0
Lz3 = 1.0
m3 = 3
eps3 = 1e-8
w3 = a2 - a1
alpha3 = np.pi / w3
def phi3_exact(x, y, z):
r = np.sqrt(x**2 + y**2)
theta = np.arctan2(y, x)
s = alpha3 * (r - a1)
return np.sin(s) * np.sin(m3 * theta)
def rho3_exact(x, y, z):
r = np.sqrt(x**2 + y**2)
theta = np.arctan2(y, x)
s = alpha3 * (r - a1)
sin_s = np.sin(s)
cos_s = np.cos(s)
sin_mt = np.sin(m3 * theta)
# -Delta(phi) + eps*phi for phi(r,theta)=sin(s)sin(m theta)
term = (alpha3**2) * sin_s - (alpha3 / r) * cos_s + (m3**2 / r**2) * sin_s + eps3 * sin_s
return term * sin_mt
model3 = Poisson()
model3.propagators.poisson.options = model3.propagators.poisson.Options(
rho=model3.em_fields.source,
stab_eps=eps3,
)
model3.em_fields.source.add_perturbation(
GenericPerturbation(rho3_exact, given_in_basis="physical")
)
domain3 = domains.HollowCylinder(a1=a1, a2=a2, Lz=Lz3)
grid3 = grids.TensorProductGrid(num_elements=(36, 96, 1))
derham_opts3 = DerhamOptions(
degree=(3, 3, 1),
bcs=(("dirichlet", "dirichlet"), None, None),
)
sim3 = Simulation(
model=model3,
domain=domain3,
grid=grid3,
derham_opts=derham_opts3,
verbose=False,
)
sim3.run(one_time_step=True, verbose=False)
sim3.pproc(verbose=False)
sim3.load_plotting_data(verbose=False)
Starting simulation run for model Poisson ...
Struphy run finished.
100%|██████████| 2/2 [00:00<00:00, 1078.50it/s]
100%|██████████| 2/2 [00:01<00:00, 1.22it/s]
100%|██████████| 2/2 [00:00<00:00, 135.15it/s]
[8]:
# Annulus diagnostics and plots in physical coordinates only
t3_last = max(sim3.spline_values.em_fields.phi_log.data.keys())
X3 = sim3.grids_phy[0][:, :, 0]
Y3 = sim3.grids_phy[1][:, :, 0]
phi3_num = sim3.spline_values.em_fields.phi_log.data[t3_last][0][:, :, 0]
phi3_ref = phi3_exact(X3, Y3, 0.0)
err3 = phi3_num - phi3_ref
err3_max = np.max(np.abs(err3))
fig3, axs3 = plt.subplots(1, 3, figsize=(15, 4.8), constrained_layout=True)
im30 = axs3[0].contourf(X3, Y3, phi3_num, levels=40, cmap="viridis")
axs3[0].set_title("Numerical $\\phi_h(x,y)$ on annulus")
axs3[0].set_xlabel("x")
axs3[0].set_ylabel("y")
axs3[0].set_aspect("equal")
plt.colorbar(im30, ax=axs3[0])
im31 = axs3[1].contourf(X3, Y3, phi3_ref, levels=40, cmap="viridis")
axs3[1].set_title("Analytic $\\phi_{\\mathrm{exact}}(x,y)$ on annulus")
axs3[1].set_xlabel("x")
axs3[1].set_ylabel("y")
axs3[1].set_aspect("equal")
plt.colorbar(im31, ax=axs3[1])
im32 = axs3[2].contourf(X3, Y3, err3, levels=40, cmap="coolwarm")
axs3[2].set_title("Error $\\phi_h-\\phi_{\\mathrm{exact}}$")
axs3[2].set_xlabel("x")
axs3[2].set_ylabel("y")
axs3[2].set_aspect("equal")
plt.colorbar(im32, ax=axs3[2])
print(f"Annulus max-norm error ||phi_h - phi_exact||_inf = {err3_max:.3e}")
Annulus max-norm error ||phi_h - phi_exact||_inf = 1.101e-05